Hierarchical Clustering
For hierarchical clustering we will use the iris dataset. To make the clustering dendrogram less crowded we will select 30 instances from the iris dataset.
indices <- sample(1:nrow(iris), 30)
dataset <- iris[indices, -5]To do hierachical clustering we will use the hclust method:
hc <- hclust(dist(dataset), method="ave")We have different options to use in the method argument to define the distance between the clusters as:
- the distance between the closests points of the clusters (
single) - the distance between the farthests points of the clusters (
complete) - the average distance between all points in the clusters (
average) - the distance between the centroids of the clusters (
centroid) - and others (see
?hclustfor a list)
To plot the dendrogram and cut the tree into three clusters:
# Plot the dendrogram
plot(hc, hang = -1, labels=iris$Species[indices])
# cut tree into 3 clusters
rect.hclust(hc, k=3)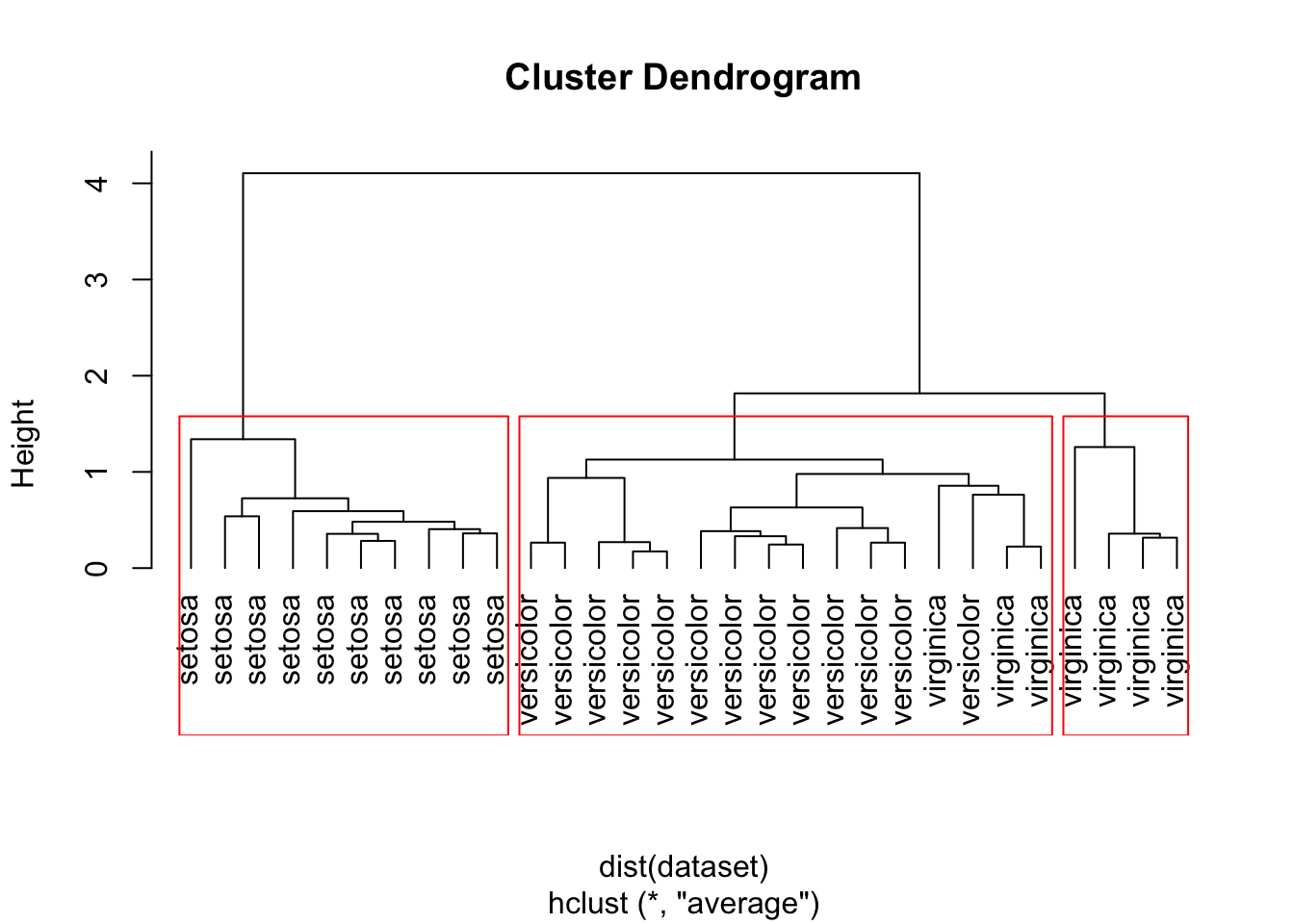
# groups contain the distribution of the samples to the clusters/groups
groups <- cutree(hc, k=3)Using single
hc <- hclust(dist(dataset), method="single")
# Plot the dendrogram
plot(hc, hang = -1, labels=iris$Species[indices])
# cut tree into 3 clusters
rect.hclust(hc, k=3)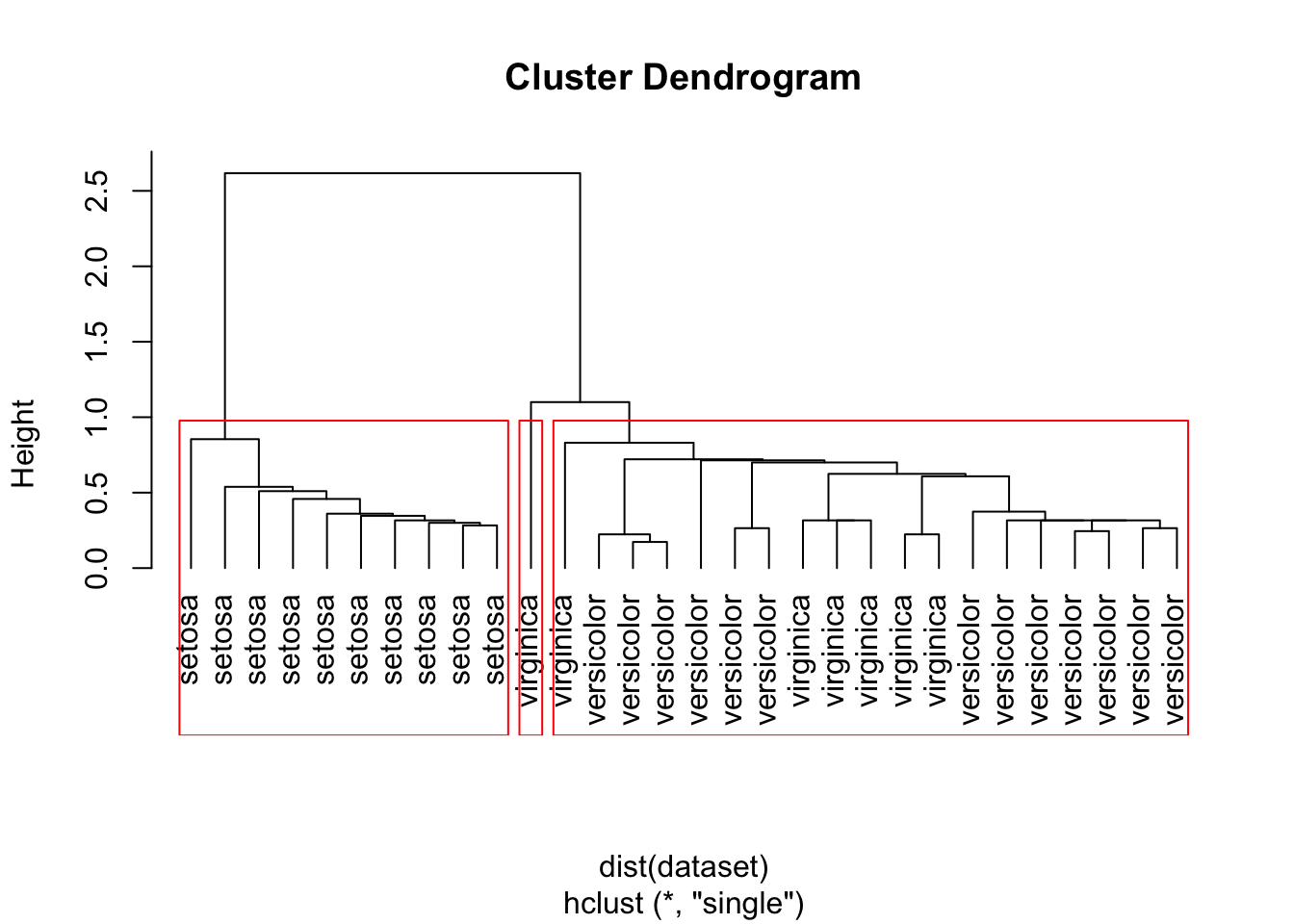
# groups contain the distribution of the samples to the clusters/groups
groups <- cutree(hc, k=3)